Note
Go to the end to download the full example code
Rotated Maximum Covariance Analysis#
Rotated Maximum Covariance Analysis (MCA) between two data sets.
# Load packages and data:
import numpy as np
import xarray as xr
import matplotlib.pyplot as plt
from matplotlib.gridspec import GridSpec
from cartopy.crs import Orthographic, PlateCarree
from cartopy.feature import LAND
from xeofs.models import MCA, MCARotator
Create 2 different DataArrays
t2m = xr.tutorial.load_dataset("air_temperature")["air"]
da1 = t2m.isel(lon=slice(0, 26))
da2 = t2m.isel(lon=slice(27, None))
Perform MCA
mca = MCA(n_modes=20, standardize=False, use_coslat=True)
mca.fit(da1, da2, dim="time")
<xeofs.models.mca.MCA object at 0x7fa6f9945b10>
Apply Varimax-rotation to MCA solution
rot = MCARotator(n_modes=10)
rot.fit(mca)
<xeofs.models.mca_rotator.MCARotator object at 0x7fa6f935a610>
Get rotated singular vectors, projections (PCs), homogeneous and heterogeneous patterns:
singular_vectors = rot.components()
scores = rot.scores()
hom_pats, pvals_hom = rot.homogeneous_patterns()
het_pats, pvals_het = rot.heterogeneous_patterns()
When two fields are expected, the output of the above methods is a list of
length 2, with the first and second entry containing the relevant object for
X and Y. For example, the p-values obtained from the two-sided t-test
for the homogeneous patterns of X are:
pvals_hom[0]
Create a mask to identifiy where p-values are below 0.05
hom_mask = [values < 0.05 for values in pvals_hom]
het_mask = [values < 0.05 for values in pvals_het]
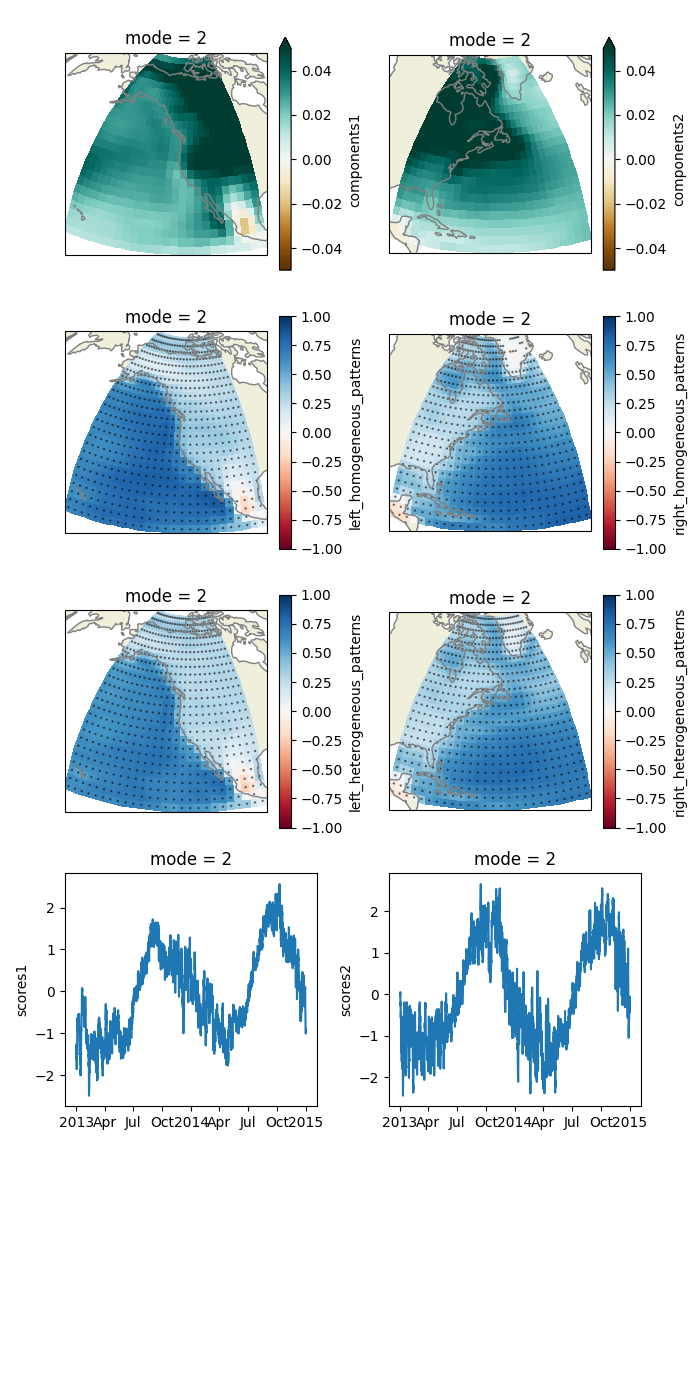
Plot some relevant quantities of mode 2.
lonlats = [
np.meshgrid(pvals_hom[0].lon.values, pvals_hom[0].lat.values),
np.meshgrid(pvals_hom[1].lon.values, pvals_hom[1].lat.values),
]
proj = [
Orthographic(central_latitude=30, central_longitude=-120),
Orthographic(central_latitude=30, central_longitude=-60),
]
kwargs1 = {"cmap": "BrBG", "vmin": -0.05, "vmax": 0.05, "transform": PlateCarree()}
kwargs2 = {"cmap": "RdBu", "vmin": -1, "vmax": 1, "transform": PlateCarree()}
mode = 2
fig = plt.figure(figsize=(7, 14))
gs = GridSpec(5, 2)
ax1 = [fig.add_subplot(gs[0, i], projection=proj[i]) for i in range(2)]
ax2 = [fig.add_subplot(gs[1, i], projection=proj[i]) for i in range(2)]
ax3 = [fig.add_subplot(gs[2, i], projection=proj[i]) for i in range(2)]
ax4 = [fig.add_subplot(gs[3, i]) for i in range(2)]
for i, a in enumerate(ax1):
singular_vectors[i].sel(mode=mode).plot(ax=a, **kwargs1)
for i, a in enumerate(ax2):
hom_pats[i].sel(mode=mode).plot(ax=a, **kwargs2)
a.scatter(
lonlats[i][0],
lonlats[i][1],
hom_mask[i].sel(mode=mode).values * 0.5,
color="k",
alpha=0.5,
transform=PlateCarree(),
)
for i, a in enumerate(ax3):
het_pats[i].sel(mode=mode).plot(ax=a, **kwargs2)
a.scatter(
lonlats[i][0],
lonlats[i][1],
het_mask[i].sel(mode=mode).values * 0.5,
color="k",
alpha=0.5,
transform=PlateCarree(),
)
for i, a in enumerate(ax4):
scores[i].sel(mode=mode).plot(ax=a)
a.set_xlabel("")
for a in np.ravel([ax1, ax2, ax3]):
a.coastlines(color=".5")
a.add_feature(LAND)
plt.tight_layout()
plt.savefig("rotated_mca.jpg")
Total running time of the script: (0 minutes 2.207 seconds)
