Note
Go to the end to download the full example code
Maximum Covariance Analysis#
Maximum Covariance Analysis (MCA) between two data sets.
# Load packages and data:
import numpy as np
import xarray as xr
import matplotlib.pyplot as plt
from matplotlib.gridspec import GridSpec
from cartopy.crs import Orthographic, PlateCarree
from cartopy.feature import LAND
from xeofs.models import MCA
Create 2 different DataArrays
t2m = xr.tutorial.load_dataset("air_temperature")["air"]
da1 = t2m.isel(lon=slice(0, 26))
da2 = t2m.isel(lon=slice(27, None))
Perform MCA
mca = MCA(n_modes=20, standardize=False, use_coslat=True)
mca.fit(da1, da2, dim="time")
<xeofs.models.mca.MCA object at 0x7fa6f9db1d50>
Get singular vectors, projections (PCs), homogeneous and heterogeneous patterns:
singular_vectors = mca.components()
scores = mca.scores()
hom_pats, pvals_hom = mca.homogeneous_patterns()
het_pats, pvals_het = mca.heterogeneous_patterns()
When two fields are expected, the output of the above methods is a list of
length 2, with the first and second entry containing the relevant object for
X and Y. For example, the p-values obtained from the two-sided t-test
for the homogeneous patterns of X are:
pvals_hom[0]
Create a mask to identifiy where p-values are below 0.05
hom_mask = [values < 0.05 for values in pvals_hom]
het_mask = [values < 0.05 for values in pvals_het]
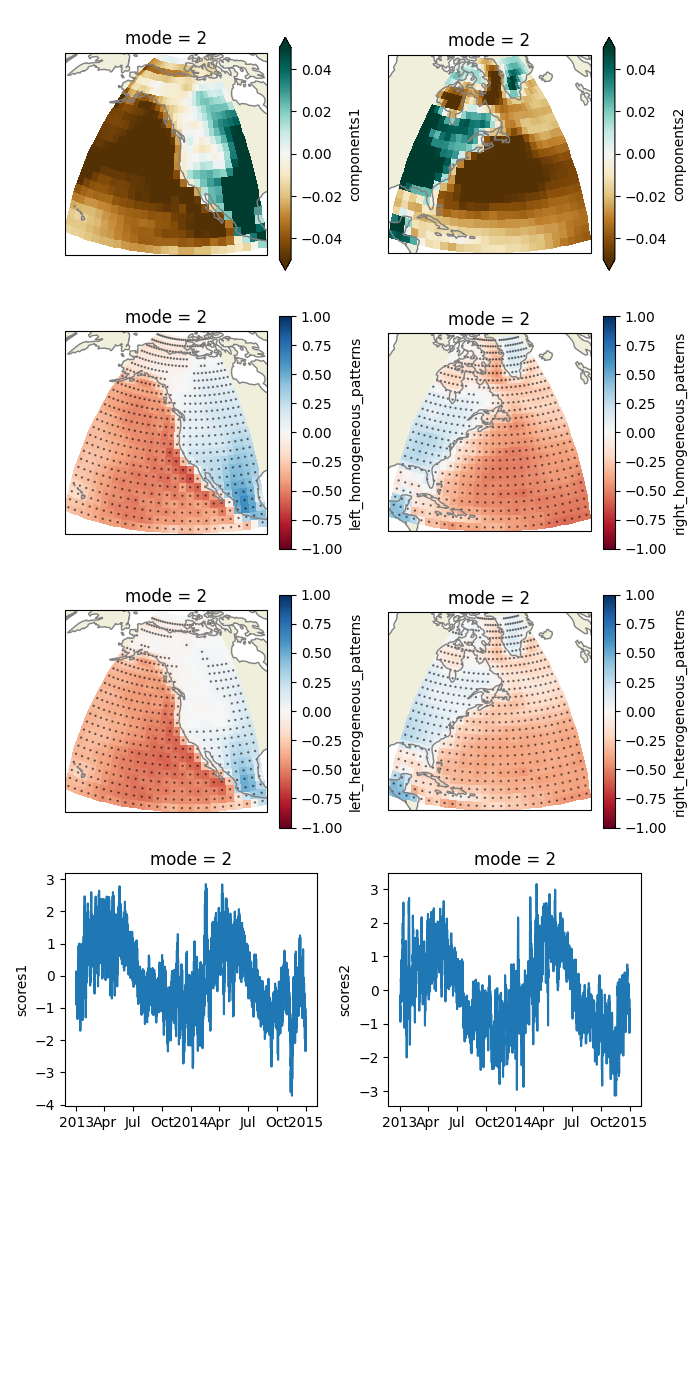
Plot some relevant quantities of mode 2.
lonlats = [
np.meshgrid(pvals_hom[0].lon.values, pvals_hom[0].lat.values),
np.meshgrid(pvals_hom[1].lon.values, pvals_hom[1].lat.values),
]
proj = [
Orthographic(central_latitude=30, central_longitude=-120),
Orthographic(central_latitude=30, central_longitude=-60),
]
kwargs1 = {"cmap": "BrBG", "vmin": -0.05, "vmax": 0.05, "transform": PlateCarree()}
kwargs2 = {"cmap": "RdBu", "vmin": -1, "vmax": 1, "transform": PlateCarree()}
mode = 2
fig = plt.figure(figsize=(7, 14))
gs = GridSpec(5, 2)
ax1 = [fig.add_subplot(gs[0, i], projection=proj[i]) for i in range(2)]
ax2 = [fig.add_subplot(gs[1, i], projection=proj[i]) for i in range(2)]
ax3 = [fig.add_subplot(gs[2, i], projection=proj[i]) for i in range(2)]
ax4 = [fig.add_subplot(gs[3, i]) for i in range(2)]
for i, a in enumerate(ax1):
singular_vectors[i].sel(mode=mode).plot(ax=a, **kwargs1)
for i, a in enumerate(ax2):
hom_pats[i].sel(mode=mode).plot(ax=a, **kwargs2)
a.scatter(
lonlats[i][0],
lonlats[i][1],
hom_mask[i].sel(mode=mode).values * 0.5,
color="k",
alpha=0.5,
transform=PlateCarree(),
)
for i, a in enumerate(ax3):
het_pats[i].sel(mode=mode).plot(ax=a, **kwargs2)
a.scatter(
lonlats[i][0],
lonlats[i][1],
het_mask[i].sel(mode=mode).values * 0.5,
color="k",
alpha=0.5,
transform=PlateCarree(),
)
for i, a in enumerate(ax4):
scores[i].sel(mode=mode).plot(ax=a)
a.set_xlabel("")
for a in np.ravel([ax1, ax2, ax3]):
a.coastlines(color=".5")
a.add_feature(LAND)
plt.tight_layout()
plt.savefig("mca.jpg")
/home/slevang/miniconda3/envs/xeofs-docs/lib/python3.11/site-packages/cartopy/io/__init__.py:241: DownloadWarning: Downloading: https://naturalearth.s3.amazonaws.com/110m_physical/ne_110m_land.zip
warnings.warn(f'Downloading: {url}', DownloadWarning)
Total running time of the script: (0 minutes 3.519 seconds)
